Technical Reference: RLDM Classes and Methods
Wolfgang Scherrer and Bernd Funovits
2026-01-31
2_technical_reference.RmdThis document is a reference guide for RLDM classes, mathematical foundations, and estimation methods. For practical examples, see the “Getting Started” and “Case Study” vignettes.
Preliminaries
Notation
The following notation is used throughout RLDM:
| Symbol | Meaning |
|---|---|
| Dimension of the output process | |
| Dimension of the noise process , typically | |
| Dimension of the state process (state space models only) | |
Sample size, denoted n.obs in code |
|
| Expectation operator | |
| Autocovariance at lag | |
| Noise covariance matrix |
Sign Convention
RLDM uses a non-standard sign convention for AR coefficients to maintain consistency with matrix fraction descriptions.
Standard form (often seen in time series textbooks):
RLDM form (matrix fraction description):
Notice the opposite signs on the AR coefficients. This ensures consistency when working with rational matrix fractions where:
with (identity matrix).
Model Classes
Overview: Model Representations
RLDM supports three equivalent ways to represent rational linear dynamic models:
1. Left Matrix Fraction Description (LMFD):
armamod
The ARMA/VARMA representation:
where and are polynomial matrices in the lag operator .
VARMA/ARMA Models: armamod Class
The armamod class represents VARMA
(multivariate ARMA) processes in left matrix fraction form.
Structure
An armamod object is an S3 list with the following
slots:
| Slot | Type | Description |
|---|---|---|
sys |
lmfd [m,n] |
Left matrix fraction: filter |
sigma_L |
double [n,n] |
Left square root of noise covariance () |
names |
character [m] |
Component names of |
label |
character |
Descriptive label for the process |
Class attribute:
c("armamod", "rldm")
Constructor
armamod(sys, sigma_L, names = NULL, label = NULL)Example:
# Create a simple AR(1) model: y_t = 0.8*y_{t-1} + u_t
# In RLDM convention: y_t - 0.8*y_{t-1} = u_t
a_coef <- matrix(-0.8) # negative sign for standard convention
b_coef <- matrix(1)
sys <- lmfd(a_coef, b_coef)
model <- armamod(sys, sigma_L = matrix(1), names = "Output", label = "AR(1)")
model
#> AR(1): ARMA model [1,1] with orders p = 0 and q = 0
#> AR polynomial a(z):
#> z^0 [,1]
#> [1,] -0.8
#> MA polynomial b(z):
#> z^0 [,1]
#> [1,] 1
#> Left square root of noise covariance Sigma:
#> u[1]
#> u[1] 1Available Methods
methods(class = 'armamod')
#> [1] as.stspmod autocov freqresp impresp ll poles
#> [7] predict print sim spectrald str zeroes
#> see '?methods' for accessing help and source codeKey methods: - autocov() - Compute
autocovariance function - impresp() -
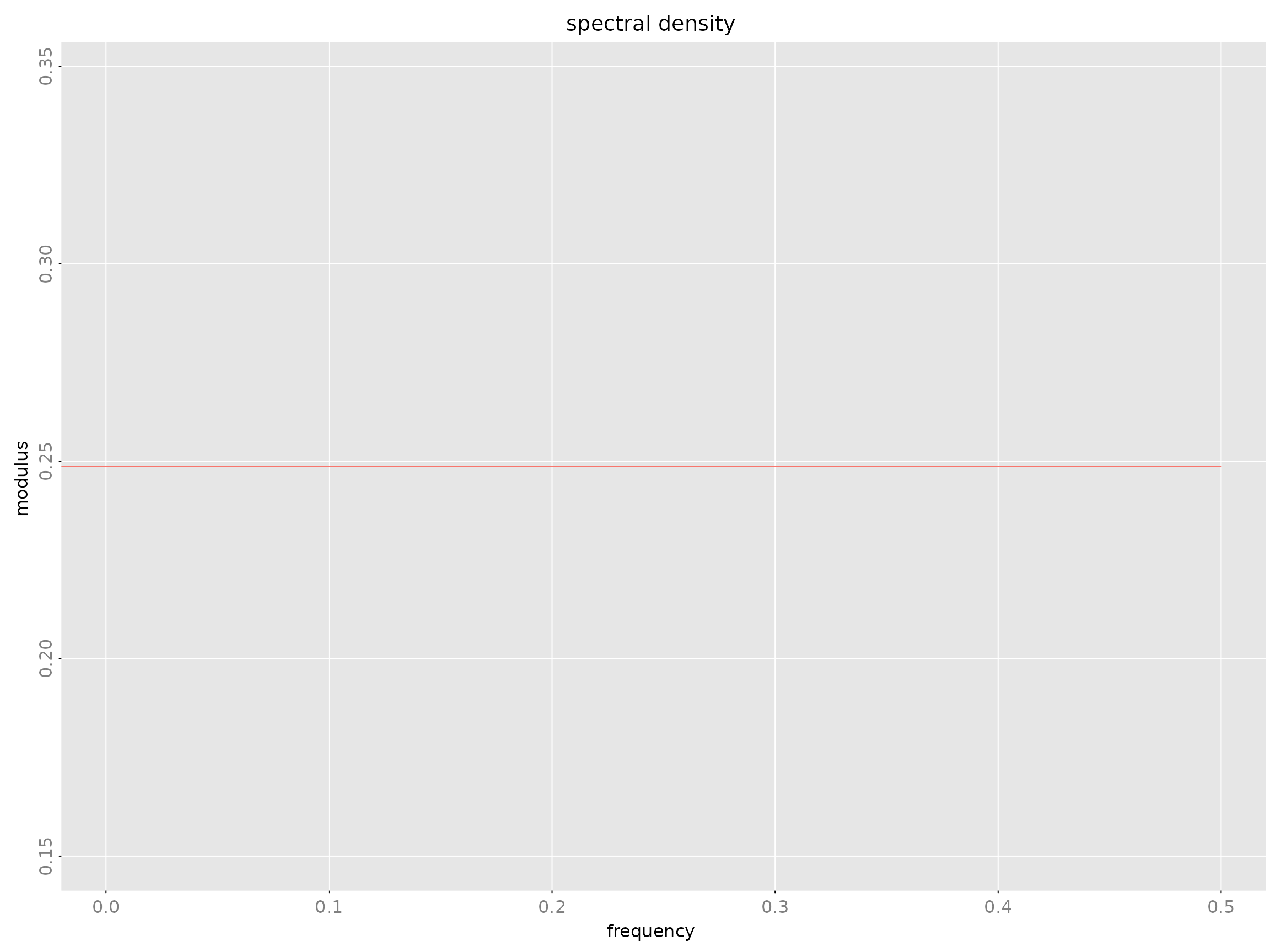
Impulse response functions - spectrald() -
Spectral density - freqresp() - Frequency
response - predict() - Forecasting -
solve_de() - Simulate from the model -
print() / plot() -
Visualization
State Space Models: stspmod Class
The stspmod class represents processes in canonical
state space form.
State space models offer several advantages over other representations:
- Minimality: The state dimension represents the essential “memory” of the system, providing the most compact representation.
-
Balancing: Realizations can be balanced to improve
numerical stability (
balanceparameter in template functions). - Latent state interpretation: Hidden states can represent meaningful unobserved components (trends, cycles, factors).
- Flexibility: Can model complex dynamics with relatively few parameters through appropriate state dimension selection.
Structure
An stspmod object is an S3 list with slots:
| Slot | Type | Description |
|---|---|---|
sys |
stsp [m,n] |
State space realization |
sigma_L |
double [n,n] |
Left factor of noise covariance |
names |
character [m] |
Component names |
label |
character |
Descriptive label |
Class attribute:
c("stspmod", "rldm")
Constructor
stspmod(sys, sigma_L, names = NULL, label = NULL)Example:
# Create a simple state space model
# s_{t+1} = 0.8*s_t + u_t
# y_t = s_t
A_matrix <- matrix(0.8)
B_matrix <- matrix(1)
C_matrix <- matrix(1)
D_matrix <- matrix(0)
sys <- stsp(A_matrix, B_matrix, C_matrix, D_matrix)
model_ss <- stspmod(sys, sigma_L = matrix(1))
model_ss
#> state space model [1,1] with s = 1 states
#> s[1] u[1]
#> s[1] 0.8 1
#> x[1] 1.0 0
#> Left square root of noise covariance Sigma:
#> u[1]
#> u[1] 1Available Methods
methods(class = 'stspmod')
#> [1] autocov freqresp impresp ll_pfilter ll pfilter
#> [7] poles predict print sim spectrald str
#> [13] zeroes
#> see '?methods' for accessing help and source codeSimilar to armamod, state space models support: -
Autocovariance, impulse responses, spectral density computation -
Forecasting and simulation - Conversion to impulse response
representation
State Space Parameterizations
RLDM supports several standard state space canonical forms through template functions:
-
tmpl_DDLC()- Diagonal-Direct-Lead-Coefficient form -
tmpl_stsp_echelon()- Echelon canonical form -
tmpl_stsp_innovation()- Innovation form
Right Matrix Fraction Description: rmfdmod Class
The rmfdmod class represents processes in right matrix
fraction form (experimental, limited methods):
rmfdmod(sys, sigma_L, names = NULL, label = NULL)Most estimation and analysis methods are not yet implemented for this class.
Derived Objects
These classes represent computed properties of process models, not model specifications themselves.
Autocovariance: autocov Class
Stores autocovariances, autocorrelations, or partial autocorrelations.
Structure
| Slot | Type | Description |
|---|---|---|
acf |
array | Correlations/covariances at each lag |
type |
character |
“covariance”, “correlation”, or “partial” |
gamma |
array [m,m,lag.max+1] |
3D array of autocovariances |
names |
character [m] |
Variable names |
label |
character |
Descriptive label |
Computation
# From data
autocov.default(obj, lag.max = 20, type = "correlation")
# From model
autocov.armamod(obj, lag.max = 20, type = "correlation")
autocov.stspmod(obj, lag.max = 20, type = "correlation")Example:
# Generate AR model and compute ACF
y <- sim(model, n.obs = 500)
acov <- autocov(model, lag.max = 15, type = "correlation")
plot(acov)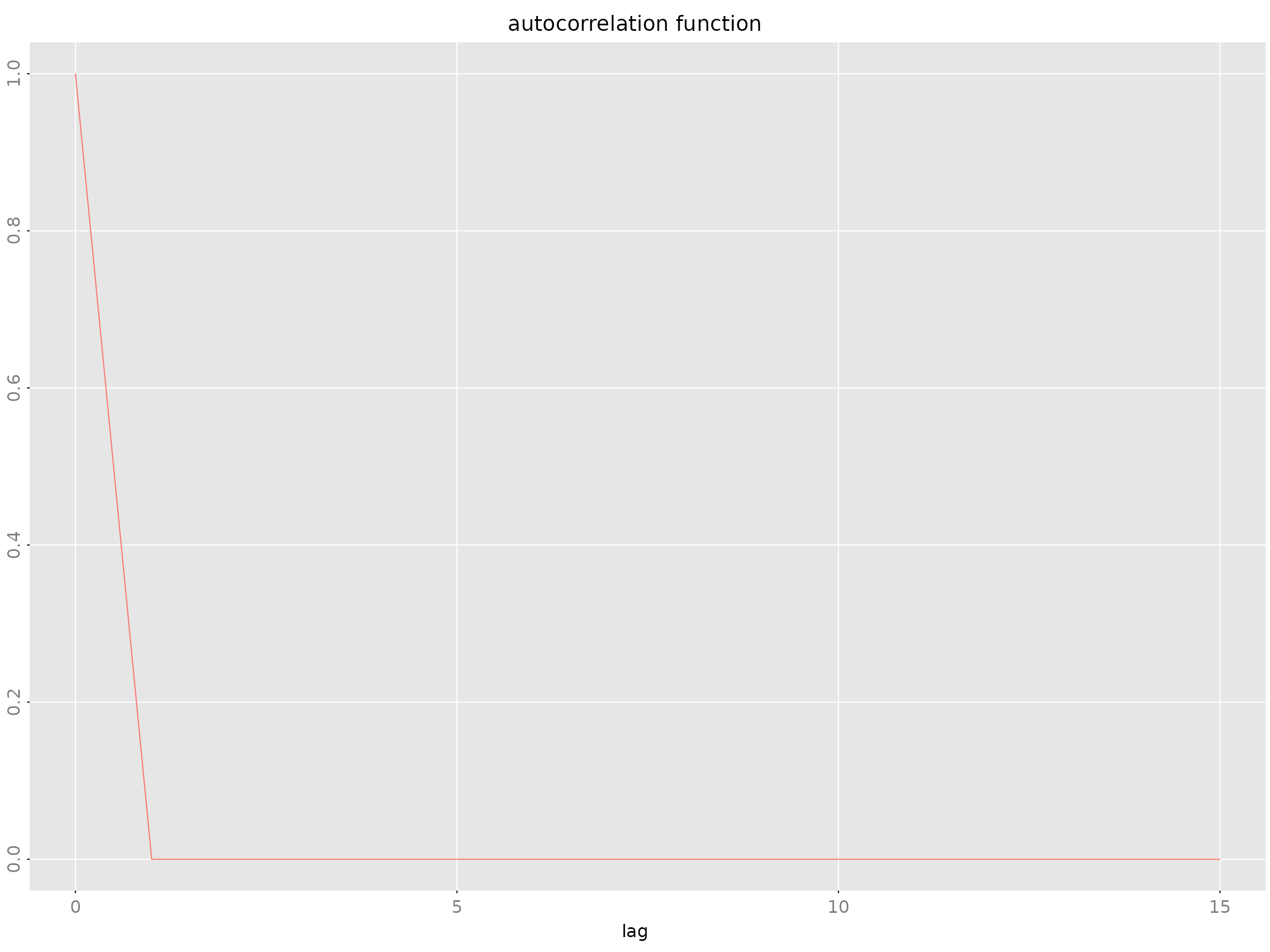
Impulse Response: impresp Class
Stores impulse response coefficients: where .
Structure
| Slot | Type | Description |
|---|---|---|
irf |
array [m,n,n.lags] |
IRF coefficients |
sigma_L |
double [n,n] |
Noise covariance factor |
names |
character [m] |
Output variable names |
label |
character |
Descriptive label |
Computation
impresp.armamod(obj, lag.max = 40, H = NULL)
impresp.stspmod(obj, lag.max = 40, H = NULL)Example:
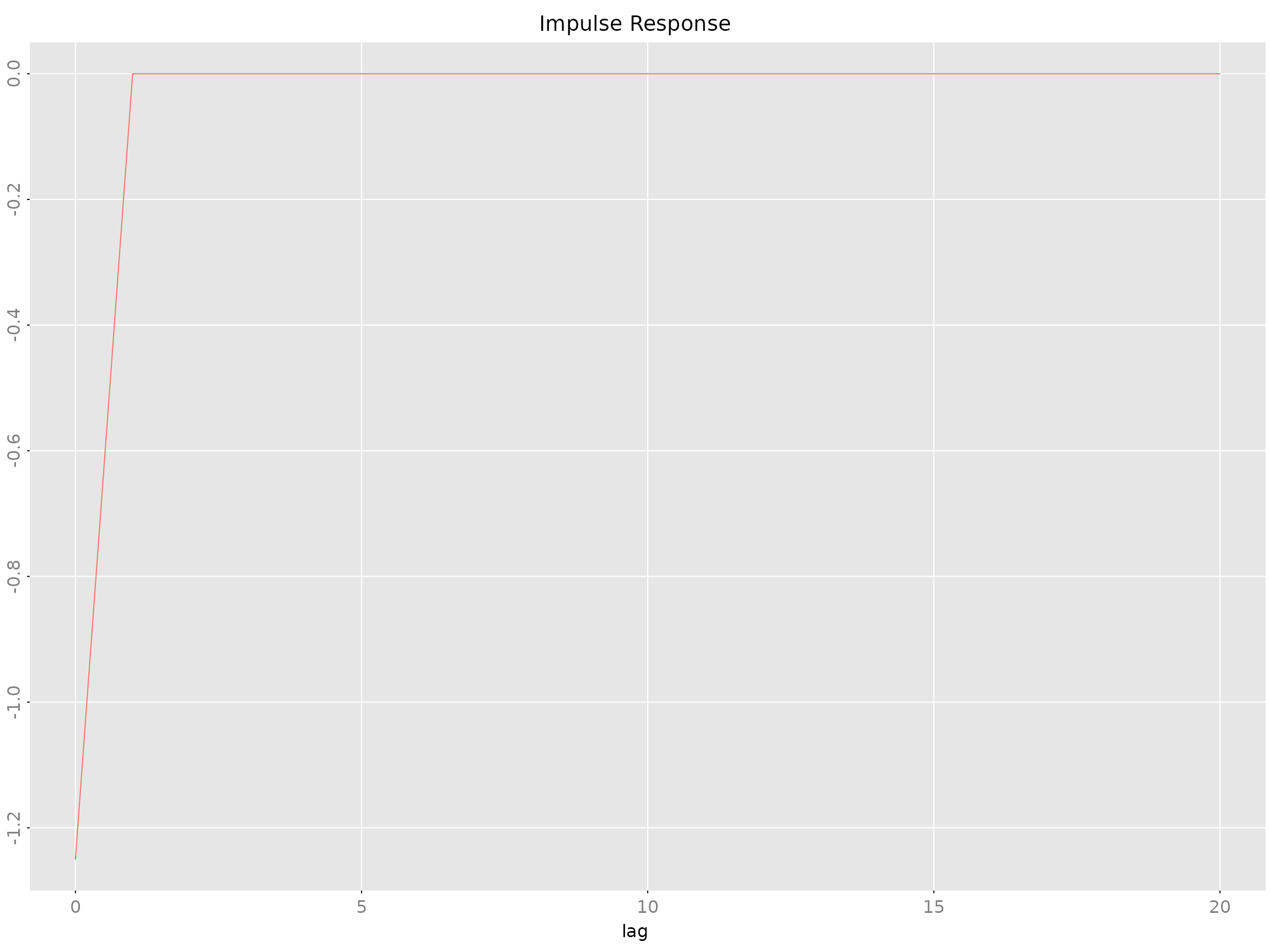
The H parameter allows custom orthogonalization
(default: Cholesky).
Spectral Density: spectrald Class
Frequency-domain representation: where is the frequency response.
Frequency Response: freqresp Class
The frequency transfer function .
freqresp.armamod(obj, n.f = 256)Forecast Error Variance Decomposition: fevardec
Class
Decomposes forecast error variance into contributions from each shock.
| Slot | Type | Description |
|---|---|---|
vd |
array [m,m,h] |
Variance decomposition |
v |
matrix [m,h] |
Forecast error variance |
names |
character [m] |
Variable names |
fevardec(obj, h.max = 40)Estimation Methods
AR Model Estimation
Yule-Walker Method: est_ar_yw()
Theory: Solves the Yule-Walker equations using Cholesky decomposition:
Advantages: - Statistically efficient (minimum variance) - Produces all AR orders simultaneously - Automatic log-determinant sequence for model selection
Example:
est_ar_yw(gamma, p.max = 10, penalty = -1)Durbin-Levinson-Whittle Method: est_ar_dlw()
Theory: Recursive algorithm for AR parameter and partial autocorrelation computation.
Advantages: - Numerically stable - Computes partial ACF - Efficient recursions
Example:
est_ar_dlw(gamma, p.max = 10, penalty = -1)Wrapper Function: est_ar()
Primary interface for AR estimation with automatic order selection:
est_ar(data, p.max = NULL, method = "auto",
ic = "AIC", mean_estimate = "sample.mean")Parameters: - ic: Information criterion
(“AIC”, “BIC”, “HQ”) - mean_estimate: How to estimate mean
(“sample.mean”, “zero”, or a vector) - method: “yw”
(Yule-Walker), “dlw” (Durbin-Levinson-Whittle), or “auto”
Returns: List with components: - model:
Estimated armamod object - p: Selected order -
stats: Criterion values for each order - ll:
Log-likelihood values
ARMA Model Estimation
Hannan-Rissanen-Kavalieris (HRK): est_arma_hrk()
Three-stage procedure for VARMA estimation without nonlinear optimization:
est_arma_hrk(data, tmpl = NULL, p = NULL, q = NULL,
mean_estimate = "zero")Stage 1: Estimate long AR model Stage 2: Compute initial ARMA estimates Stage 3: Refine using feasible GLS
Advantages: - Provides consistent initial estimates - No nonlinear optimization required - Suitable for maximum likelihood refinement
When to use: - Initial estimates for multivariate ARMA - Moderate to high-dimensional systems - When you want to avoid local optima
State Space Estimation
CCA Method: est_stsp_ss(method = "cca")
Canonical Correlation Analysis for determining state space order and initial estimates.
est_stsp_ss(data, method = "cca", s = NULL,
keep_models = FALSE, ...)Advantages: - Data-driven order selection - Suitable when no prior knowledge of system structure - Good numerical properties
State Dimension Selection: When
s = NULL (default), CCA automatically selects state
dimension using singular value criteria. For manual selection, consider:
- Singular Value Criterion (SVC): Look for a gap in
singular values of the weighted Hankel matrix - Information
Criteria: AIC/BIC computed for different state dimensions -
Cross-validation: Out-of-sample prediction performance
- Interpretability: Choose dimension that matches
expected number of latent factors
In practice, start with automatic selection via
est_stsp_ss() with s = NULL, then inspect the
singular values plot (if available) or compare information criteria
across dimensions.
Maximum Likelihood Estimation
General ML Framework: ll_FUN()
Create likelihood functions for structured parameter templates:
llfun <- ll_FUN(tmpl, data, skip = 0, which = "concentrated")Then optimize using standard R optimizers:
Parameter Templates
Templates map deep model parameters to linear parameters for optimization:
-
tmpl_DDLC()- Diagonal-Direct-Lead-Coefficient -
tmpl_arma_echelon()- ARMA in echelon form -
tmpl_stsp_echelon()- State space in echelon form
Workflow:
# Create template from initial estimate
tmpl <- tmpl_DDLC(model_initial, sigma_L = 'identity')
# Extract starting parameter vector
theta0 <- extract_theta(model_initial, tmpl)
# Create and optimize likelihood
llfun <- ll_FUN(tmpl, data, which = "concentrated")
opt <- optim(theta0, llfun, method = "BFGS", control = list(fnscale = -1, maxit = 500))
# Reconstruct model from optimized parameters
model_ml <- fill_template(opt$par, tmpl)Particle Filtering
Overview
Particle filters (Sequential Monte Carlo methods) extend Kalman filtering to nonlinear and/or non-Gaussian state space models. While the Kalman filter provides optimal closed-form solutions for linear Gaussian models, particle filters use Monte Carlo approximation to handle:
- Nonlinear state transitions:
- Non-Gaussian observations: with arbitrary distribution
- Complex noise structures
The RLDM implementation includes three particle filter variants:
- SIR (Bootstrap) filter: Basic sampling importance resampling
- APF (Auxiliary Particle Filter): Two-stage resampling with lookahead
- Optimal proposal: For linear Gaussian models, approaches Kalman filter performance
Basic Usage: Linear Gaussian Comparison
For linear Gaussian models, particle filters approximate the Kalman filter solution:
library(RLDM)
# Create a linear Gaussian state space model
m <- 1; s <- 2; n <- s
tmpl <- tmpl_stsp_full(m, n, s, sigma_L = "chol")
model <- r_model(tmpl, bpoles = 1, sd = 0.5)
data <- sim(model, n.obs = 100)
y <- data$y
# Kalman filter (optimal for linear Gaussian)
kf_result <- kf(model, y)
# Particle filters
pf_sir <- pfilter(model, y, N_particles = 1000, method = "sir")
pf_apf <- pfilter(model, y, N_particles = 1000, method = "apf")
pf_opt <- pfilter(model, y, N_particles = 1000, method = "optimal")
# Compare filtered states
plot(kf_result$a[,1], type = "l", col = "black", lwd = 2,
main = "Filtered State Estimates")
lines(pf_sir$filtered_states[,1], col = "blue", lty = 2)
lines(pf_apf$filtered_states[,1], col = "red", lty = 2)
lines(pf_opt$filtered_states[,1], col = "green", lty = 2)
legend("topright", legend = c("Kalman", "SIR", "APF", "Optimal"),
col = c("black", "blue", "red", "green"), lty = c(1, 2, 2, 2), lwd = c(2, 1, 1, 1))Likelihood Approximation
Particle filters can approximate the log-likelihood for nonlinear/non-Gaussian models where exact Kalman filter likelihood is unavailable:
# Kalman filter likelihood (exact for linear Gaussian)
ll_kf <- ll_kf(model, y)
# Particle filter likelihood approximation (averaged over multiple runs)
ll_pf <- ll_pfilter(model, y, N_particles = 1000, filter_type = "apf", N_runs = 10)
cat("Kalman filter LL:", ll_kf, "\n")
cat("Particle filter LL:", ll_pf, "\n")
cat("Difference:", ll_pf - ll_kf, "\n")Nonlinear and Non-Gaussian Examples
RLDM includes example scripts demonstrating particle filter superiority for nonlinear/non-Gaussian models:
-
inst/examples/nonlinear_sin.R- Nonlinear state transition: -
inst/examples/nonlinear_poisson.R- Non-Gaussian observation: -
inst/examples/stochastic_volatility.R- Stochastic volatility model:
Each example compares particle filter performance with approximate Kalman filter variants (Extended Kalman Filter, Gaussian approximation, log-squared transformation).
Diagnostics and Visualization
The plot.pfilter() method provides several diagnostic
plots:
pf <- pfilter(model, y, N_particles = 500, method = "sir")
# Filtered state estimates
plot(pf, type = "states")
# Effective Sample Size (ESS) over time
plot(pf, type = "ess")
# Particle weight distribution at final time
plot(pf, type = "weights")
# Log-likelihood contributions per time step
plot(pf, type = "likelihood")Performance Considerations
- Particle count: More particles improve accuracy but increase computation (typical: 1000-10000)
- Resampling methods: Systematic (default), multinomial, or stratified
- ESS threshold: Adaptive resampling when ESS < threshold × N_particles (default: 0.5)
- Optimal proposal: Use for linear Gaussian models to minimize weight degeneracy
References
- Gordon, N. J., Salmond, D. J., & Smith, A. F. M. (1993). Novel approach to nonlinear/non-Gaussian Bayesian state estimation
- Pitt, M. K., & Shephard, N. (1999). Filtering via simulation: Auxiliary particle filters
- Doucet, A., Godsill, S., & Andrieu, C. (2000). On sequential Monte Carlo sampling methods for Bayesian filtering
Solution and Simulation
Solving Difference Equations
Forward Solution: solve_de()
Simulate forward from initial conditions:
y <- solve_de(sys, u, y_init = NULL)Inverse Solution: solve_inverse_de()
Extract residuals given data (solve backwards):
u <- solve_inverse_de(sys, y)$uMathematical Foundations
Stationary ARMA Processes
An ARMA process is described by:
where: - (AR polynomial) - (MA polynomial) - is white noise with
The process is stationary if all roots of lie outside the unit circle.
The stationary solution is:
where satisfy: .
Autocovariance Function
For stationary ARMA processes, the autocovariance function satisfies the generalized Yule-Walker equations:
Method Selection Guide
Choosing Between AR, ARMA, and State Space
| Method | When to Use | Advantages | Disadvantages |
|---|---|---|---|
| AR | Baseline, simple systems | Fast, stable, interpretable | May need high order |
| ARMA | Parsimonious fits | Fewer parameters | Parameter estimation harder |
| State Space | Complex systems, latent factors | Flexible, interpretable states | Requires order selection |
Choosing State Space Parameterizations
RLDM supports several state space parameterizations, each with different strengths:
| Parameterization | When to Use | Key Features | Estimation Method |
|---|---|---|---|
| CCA (Canonical Correlation Analysis) | Initial exploration, data-driven order selection | Automatic state dimension selection, good initial estimates | est_stsp_ss(method = "cca") |
| DDLC (Diagonal-Direct-Lead-Coefficient) | Maximum likelihood refinement, stable optimization | Numerically stable, orthogonal to equivalence classes | ML with tmpl_DDLC() + optim()
|
| Echelon Form | Parsimonious canonical representation, structural restrictions | Minimal parameters via Kronecker indices, identifiable |
tmpl_stsp_echelon() + ML |
Guidelines: 1. Start with CCA for
initial model discovery and automatic order selection 2. Refine
with DDLC for maximum likelihood estimation with good numerical
properties 3. Use echelon form when you need canonical,
identifiable representations with structural restrictions 4.
Convert between forms using pseries2stsp()
and template functions as needed
Model Comparison Workflow
-
Start with AR using
est_ar() -
Try state space with
est_stsp_ss()for dimensionality reduction - Compare with ARMA using HRK initial estimates + ML refinement
- Refine best model with parameter templates and optimization
- Validate using residual diagnostics, ACF, and spectral plots
Estimation Method Selection
| Task | Recommended Method |
|---|---|
| Order selection | AIC via est_ar()
|
| Quick state space | CCA via est_stsp_ss()
|
| Multivariate ARMA | HRK via est_arma_hrk()
|
| Maximum likelihood | Parameter templates + optim()
|
| Highly structured models | Custom templates with tmpl_*()
|
References
(Scherrer and Deistler 2019)